As hospitals and public health organizations switch to using genomic data for testing, searching through genomic data can still take some time. Y Combinator-backed startup, One Codex, wants to help researchers, clinicians and public health officials, who have sequenced more than 100,000 genomes and created petabytes of data, to search this data.
Founded by Nick Greenfield, a former data scientist, and Nik Krumm, who has a PhD in Genome Sciences from the University of Washington, One Codex is a service platform for genomics driven by the genomic sequencing revolution.
Apart from using search technology, the platform also acts as an indexed, curated reference.
One Codex, which is currently in open beta, can search its growing database of 30,000 bacteria, viruses and fungi in real time and identify data sets in minutes (millions of DNA base pairs per second).
Currently, the most commonly used tool for genome searching is by using an algorithm called BLAST, Basic Local Alignment Search Tool, which compares primary biological sequence information.
“While there are a lot of “it depends” … the number we’re comfortable with is somewhere in the 1000 to 1500 [times faster] plus range,” he said.
Uploading a file to BLAST took 2 minutes and 30 seconds to process for Greenfield, and for the One Codex system that number was less than 1/20th of a second, meaning it was upwards of 3,000 times faster in this case.
Greenfield says the company wants to bring this technology to the clinical infectious disease market.
“Instead of using a specific test for tuberculosis, the doctor would take a sample, sequence that sample and transform that biology into data, and then exhaustively search that data against all the pathogens and they’ll be able to tell you if you have TB, the type of TB and maybe this TB has antibiotic resistance,” he said.
They’re also moving into the public health and food safety sector, as agencies like the Food and Drug Administration perform half a billion food pathogen tests every year, which are now being converted to being genomics based tests.
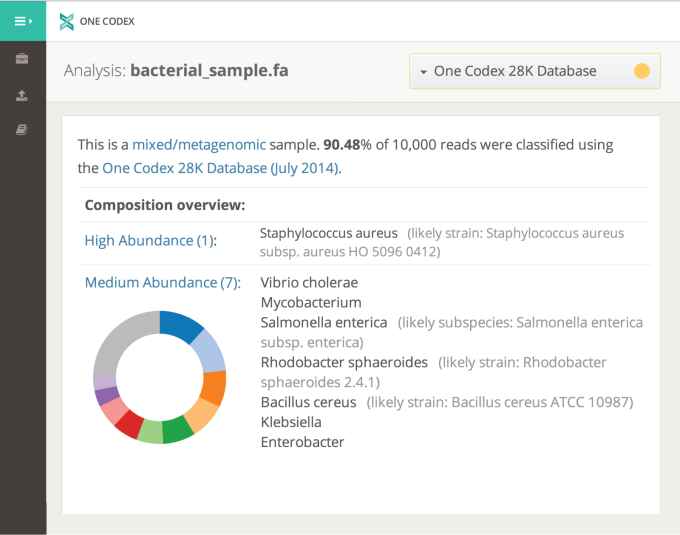
Users can upload any sequencing platform in FASTA or FASTQ format and the search platform will classify it. Both are text-based formats storing biological sequences and corresponding quality codes.
The platform uses two databases to classify user input: the RefSeq 65 Complete Genomes database, which includes 2,718 bacterial genomes and 2,318 viral genomes, and also the One Codex 28,000 database has the RefSeq 65 database as well as 22,710 additional genomes from the National Center for Biotechnology Information repository, bringing a total of 23,498 bacterial genomes, 3,995 viral genomes and 364 fungal genomes.
Right now the company is focusing on testing their platform with hospitals and agencies before implementing a way to monetize its service. Give that six years ago, it cost $10 million to sequence a human genome and today it costs roughly $1,200, these guys are at the right place at the right time.
